Samples and data sources
Available summary statistics
Summary statistics from GWAS, including only White European female samples, were used for all variables, except for sensitivity analyses conducted in White European males. Samples were selected to minimise overlap between the exposure and outcome samples, while allowing for the largest possible sample sizes. No statistical method was used to predetermine sample size. Main exposure variables of interest included estradiol levels and factors related to lifetime estradiol exposure (i.e., reproductive span, age at menarche, age at menopause, and number of childbirths). The main outcome variables of interest were brain age gap, Alzheimer’s disease, and depression.
To ensure robustness to effects of menopausal status previously shown to impact GWAS results24, several datasets were included for estradiol levels. The following samples from the UKB were included: a combined sample of pre- and postmenopausal females, a sample of only premenopausal females, and a sample of only postmenopausal females24. Main analyses in each sample were performed using GWAS of continuous estradiol levels. Sensitivity analyses were conducted in each sample using a binary approach, examining estradiol levels above and below the detection limit24 to increase sample sizes. Further, analyses on continuous estradiol levels were replicated in an independent sample of postmenopausal females from the LIFE-Adult and LIFE-Heart studies45 and a male-only sample from the UKB46.
For Alzheimer’s disease47 as an outcome, a larger GWAS for age at menopause was used that was not limited to natural menopause48 and a sensitivity analysis was conducted using age at natural menopause49. To avoid sample overlap, analyses using age at menarche50 and age at natural menopause49 as exposures and brain age gap and depression51 as outcomes were conducted using GWAS summary statistics from a smaller, independent sample. Supplementary analyses were conducted using factors related to exogenous hormone use and health-related procedures likely impacting estradiol levels, including oral contraceptive use, HRT use, history of hysterectomy, and history of oophorectomy52 as exposure variables. As the GWAS on depression51 included both single episode and recurrent depression, sensitivity analyses were conducted using recurrent depression53 as an outcome variable to check whether disease burden influenced the results found. Further, as some of the analyses using depression51 as an outcome had sample overlap, these analyses were replicated using a subsample of the depression GWAS that did not include the UKB sample53. BMI54 was included in multivariable sensitivity analyses, to account for potential pleiotropic effects16,17,21.
For estradiol levels in the independent replication sample45, estradiol levels in the male-only sample46, age at menarche48,50, age at (natural) menopause48,49, number of childbirths, oral contraceptive use, HRT use, history of hysterectomy, history of oophorectomy52, and BMI54 summary statistics were publicly available in data repositories or obtained from the IEU OpenGWAS project52. For estradiol levels in the premenopausal sample and in the postmenopausal sample24, reproductive span9, Alzheimer’s disease47 in the female-only and male-only samples, depression51, the depression subsample excluding UKB53 in the female-only and male-only samples, and recurrent depression53 summary statistics were requested and received from the respective authors. For depression in the male-only sample, a GWAS meta-analysis was run in line with the female-only GWAS51 including data from the UKB10, MoBa25, and a previously conducted GWAS, received from the authors upon request53 (see Supplementary Notes 1, 2, and 5 for details). See Table 3 and Supplementary Table 1 for a complete overview of the samples and datasets included in the present study.
UKB samples
As, to the best of our knowledge, no previously conducted sex-stratified GWAS on brain age gap existed, we ran separate GWAS on female- and male-only samples using data from the UKB. Further, to avoid sample overlap in the analyses using estradiol levels in the combined sample of pre- and postmenopausal females, we ran a female-specific GWAS using data from the UKB. The UKB complies with the Helsinki Declaration, with informed consent obtained from all participants. For the male-only GWAS on brain age gap, only genetic males (XY chromosome) with White British ancestry were included. For the female-only GWAS on brain age gap and estradiol levels, only genetic females (XX chromosome) with White British ancestry were included. For both GWAS on brain age gap, only participants with both T1-weighted and diffusion-weighted MRI data were included, and participants with ICD-10 diagnoses known to impact brain health were excluded (see Supplementary Note 1 for details on exclusions). For the GWAS on estradiol levels, participants with serum blood sample data from their first visit to the UKB assessment centre (2006 – 2010) were included. To avoid sample overlap, participants were excluded from the estradiol levels sample if they were included in the brain age gap sample. Further, participants who answered, “prefer not to answer” or “do not know/not sure” on any of the covariates included in the GWAS on continuous estradiol levels were excluded (Supplementary Note 1).
Sample overlap
Sample overlap was avoided for all analyses using Alzheimer’s disease and recurrent depression as outcomes, as well as the sensitivity analyses in the depression53 subsample, excluding UKB. Sample overlap with brain age gap and depression as outcomes was avoided for age at menarche and age at menopause as exposures by using datasets that did not include the UKB sample. Further, sample overlap with brain age gap as an outcome was avoided for estradiol levels (combined pre- and postmenopausal samples) and reproductive span as exposures by excluding the MRI sample from the UKB. However, sample overlap could not be avoided for the analyses using continuous estradiol levels in the combined pre- and postmenopausal sample and reproductive span as exposures and depression as an outcome (maximum sample overlap: 10.53% for estradiol levels and 36.91% for reproductive span). Further, there was sample overlap for continuous estradiol levels in the premenopausal sample and in the postmenopausal sample with brain age gap and depression (maximum sample overlap: 43.25% for premenopausal females with brain age gap as an outcome; 10.03% for premenopausal females with depression as an outcome; 26.31% for postmenopausal females with brain age gap as an outcome; 1.14% for postmenopausal females with depression as an outcome). For the number of childbirths, there was a maximum sample overlap of 5.70% with brain age gap as an outcome and 82.19% with depression as an outcome. Further, sample overlap could not be avoided for the supplementary analyses using factors related to exogenous hormone use and health-related procedures and the sensitivity analyses using estradiol levels in males with brain age gap and depression as outcomes, as well as using binary estradiol levels with depression as an outcome (see Supplementary Table 2).
Brain age gap prediction
Grey matter and white matter MRI features were derived from T1-weighted and diffusion-weighted images, respectively, and used for multimodal brain age prediction (see Alfaro-Almagro et al.55 and Miller et al.56 for details on MRI data acquisition and protocols in the UKB). Automated surface-based morphometry and subcortical segmentation pipelines in FreeSurfer v5.357 were used to process raw T1-weighted images. The standard set of FreeSurfer-derived subcortical and cortical summary statistics57 were further supplemented by a fine-grained cortical parcellation scheme58 to extract cortical thickness, area, and volume for 180 regions of interest per hemisphere. Outliers based on Euler numbers more than 4 SD above the mean were excluded, to avoid poor-quality data likely arising from movement59. The data from FreeSurfer was controlled for scanner site and intracranial volumes using linear models. Diffusion-weighted data were processed using an optimised diffusion pipeline60,61,62 and similarly controlled for scanner site using linear models. The diffusion-weighted data passed TBSS post-processing quality control through the YTTRIUM algorithm60,61,62. In total, 1,118 T1-weighted features and 912 diffusion-weighted features were included in the brain age prediction model.
Python (v3.7.4) was used for the brain age prediction. In line with previous studies on brain age gap in females7,8,9, brain age was computed using the XGBoost (eXtreme Gradient Boosting) regression model, based on a decision-tree ensemble algorithm63. Hyperparameters were tuned using a nested cross-validation with 5 inner folds for randomised search and 10 outer folds for validation of the model (see general model setup; see Supplementary Table 3 for model performance metrics). Brain age was estimated for each individual, and brain age gap was calculated by subtracting chronological age from the brain age prediction11. A final sample of N = 14,287 females was included in the brain age gap calculations and the subsequently conducted GWAS (age range: 45.13 – 81.83 years; mean = 63.56, SD = 7.27; see Supplementary Note 1 for details on the male brain age gap sample).
Estradiol levels
In the UKB, estradiol levels were measured using two-step competitive analysis on a Beckman Coulter Unicel Dxl 800 with a minimum detection limit of 175 pmol/L (for details see UKB Data Field 30800). A majority of the participants in the UKB have estradiol levels below this limit, likely due to the advanced age of the cohort. The GWAS on estradiol levels using the continuous approach included females with measurements above the detection limit (N = 34,697; age range: 40.16 – 70.17 years; mean = 48.30, SD = 5.96). As the estradiol levels were positively skewed, we performed an inverse rank normalisation on the data. Following previous studies23,24, a second GWAS was conducted using a binary approach which classified all females (N = 207,119) as either below (0) or above (1) the detection limit to increase the sample size (Supplementary Note 1).
GWAS procedure
The UKB v3 imputed genetic data was used, which has been genotyped, extensively quality controlled, and imputed by the UKB genetics team (see Bycroft et al.64 for details). PLINK 2.065 was used to run the GWAS. We performed standard quality check procedures, by excluding individuals with more than 10% missingness and setting the minor allele frequency (MAF) threshold at .005. We filtered out single nucleotide polymorphisms (SNPs) with more than 5% missingness as well as those out of the Hardy-Weinberg equilibrium (HWE) at p < 1 x 10−9. GWAS were run using the standard additive model of linear associations for the continuous variables (i.e., brain age gap and continuous estradiol levels) and logistic associations for the binary variable (i.e., binary estradiol levels). All analyses were controlled for the age of the participants at the time of measurement and the first 20 principal components to account for population structure. The GWAS on continuous estradiol levels was further controlled for menopausal status (premenopausal/postmenopausal), history of oral contraceptive use (yes/no), history of HRT use (yes/no), history of bilateral oophorectomy (yes/no), and history of hysterectomy (yes/no), as these variables are known to influence estradiol levels.
Post-GWAS annotations
For annotations of the GWAS results, Functional Mapping and Annotation of Genome-Wide Association Studies (FUMA) was used, applying the SNP2GENE function66. GWAS results were clumped using the 1000 Genomes projects phase 3 European dataset (1KG/Phase3) linkage disequilibrium (LD) structure with a maximum p-value of lead SNPs < 5 × 10−8, a maximum p-value cutoff < 0.05, an R2 threshold ≥ .6 to define independent significant SNPs, an R2 threshold ≥ .1 to define lead SNPs, and 250 kilo-bases (kb) as the maximum distance to merge LD blocks into a locus66. Positional mapping was performed using a maximum distance of 10 kb. Independent significant SNPs for each genomic region were identified, along with lead SNPs defined as SNPs with the smallest p-value for a genomic region. For estradiol levels, independent significant SNPs were linked to traits as reported in the GWAS Catalogue32 to facilitate investigation of potential pleiotropy of these variables used as exposures in the Mendelian randomization analyses. Using the GENE2FUNC function implemented by FUMA, the expression levels of the mapped genes were investigated for each of the GTEx v8 54 tissue types67 and hypergeometric tests were performed for gene-set enrichment analyses, testing whether genes are overrepresented in any of the pre-defined gene sets and their associations with various phenotypes66.
Two-sample Mendelian randomization analyses
Two-sample Mendelian randomization analyses were performed using R (v4.3.1). The main steps of the analyses are visualised in Fig. 4. All p-values for the IVW estimates were adjusted for multiple comparisons across all analyses (91 comparisons), including the supplementary, sensitivity, and follow-up analyses, using false discovery rate (FDR; 5%) correction68. The present study was preregistered on Open Science Framework (OSF; see Supplementary Note 8 for deviations from the preregistration).
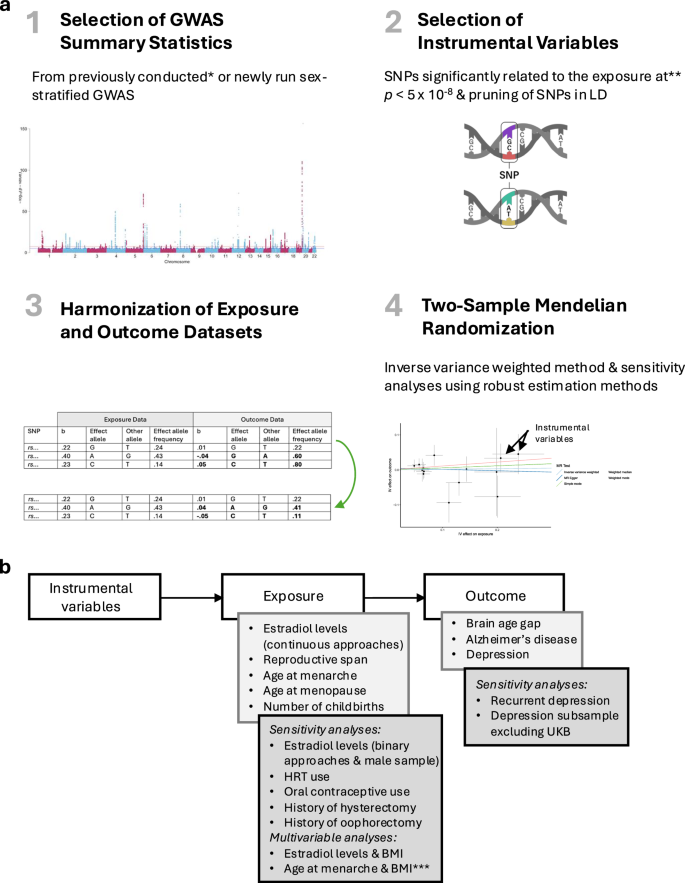
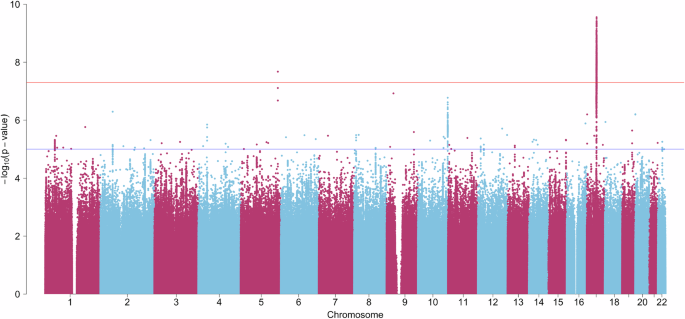
a Analysis steps. 1: Example Manhattan plot of a genome-wide association study (GWAS) for visualisation purposes. The blue/lower and red/upper lines indicate the suggestive (p < 5 × 10−5) and the genome-wide significance (p < 5 × 10−8; i.e., accounting for multiple comparisons) thresholds, respectively. The instrumental variables included in a Mendelian randomization analysis are assumed to be associated with the respective exposure variable (relevance assumption), not be associated with the outcome variable via confounding pathways (exchangeability assumption), and not affect the outcome variable in any way other than through the exposure variable (exclusion restriction assumption)15. *Requested from respective authors or openly available. 2: To satisfy the relevance assumption, the genome-wide significance threshold (p-value cut-off) was chosen for the selection of instrumental variables, when possible. **Relaxed in the case of too few available instrumental variables (see Supplementary Note 3 and Supplementary Table 4 for details). 3: Example data of exposure and outcome GWAS summary statistics for visualisation purposes. b refers to the beta-estimate of the GWAS for each single-nucleotide polymorphism (SNP). 4: Example plot of two-sample Mendelian randomization analyses for visualisation purposes. Each point represents one SNP included as an instrumental variable. Data are presented as the beta value and the confidence interval (+/− standard error) of the effect of a SNP on the exposure on the x-axis and the effect of a SNP on the outcome on the y-axis. Analyses were conducted using the inverse variance weighted method as a main estimation method (one-sided test). As the exchangeability and exclusion restriction assumptions are not verifiable15, sensitivity analyses were conducted to assess biases arising from potential violations. b Conducted analyses. ***Follow-up analysis with depression as an outcome. LD Linkage disequilibrium. HRT Hormone replacement therapy. BMI Body mass index. UKB UK Biobank.
Selection of instrumental variables
The instrumental variables included in a Mendelian randomization analysis are assumed to be associated with the respective exposure variable (relevance assumption), not be associated with the outcome variable via confounding pathways (exchangeability assumption), and not affect the outcome variable in any way other than through the exposure variable (exclusion restriction assumption)15. The relevance assumption can be verified, while the exchangeability and exclusion restriction assumptions are not verifiable15, however, a set of sensitivity analyses, including the use of robust methods and multivariable Mendelian randomization, were conducted to assess biases arising from potential violations.
To satisfy the relevance assumption, instrumental variables were selected by applying a genome-wide significance threshold of p < 5 × 10−8 (p < 5 × 10−7 or p < 5 × 10−6 in the case of few available SNPs; see Supplementary Note 3 for details on the selection of instrumental variables for each analysis) to identify SNPs associated with the respective exposure variables. Using the TwoSampleMR package (v0.5.11)69, which makes use of data from the 1000 Genomes project, significant SNPs in LD were pruned to include only the most significant SNP, and the exposure and outcome datasets were harmonised.
First-stage F-statistics were approximated for all instrumental variables as a measure of instrument strength (F < 10 indicating potential weak instrument bias), using the MendelianRandomization package (v0.9.0)70. The instrumental variables did not indicate weak instrument bias for any of the main analyses. For the sensitivity analyses, weak instrument bias was only indicated for the instruments used for the number of childbirths as an exposure with the depression53 subsample excluding UKB as an outcome (F = 0.80) and with recurrent depression as an outcome (F = 0.70; Supplementary Table 4). Furthermore, Cochran’s Q was computed as a measure of heterogeneity, which assesses whether the instrumental variables underlie the same causal parameter and can thereby be used as an indicator of potential pleiotropy (i.e., instrumental variables affect the outcome through other biological pathways) when significant15.
Univariable analyses
Univariable two-sample Mendelian randomization analyses were conducted using the GWAS summary statistics. Univariable analyses were performed between all separate exposure variables and all separate outcome variables (Fig. 4). Odds ratios were log-transformed before conducting analyses. Analyses were performed using the TwoSampleMR package (v0.5.11)69. For the main analyses, the IVW method was applied, as it maximises statistical power15. Results were compared to more robust Mendelian randomization approaches, including MR-Egger, which allows for directional pleiotropy, as well as the weighted median, simple mode, and weighted mode estimation methods, which are more robust to outliers and make varying assumptions regarding the validity of the instrumental variables (see Burgess and Thompson, 202115 for details). Using the MRlap package (v0.0.3.2)71, MRlap was performed for analyses with sample overlap. MRlap corrects for potential biases arising from sample overlap, weak instruments, and winner’s curse (i.e., the overestimation of effects in discovery GWAS)71.
Multivariable analyses
Multivariable Mendelian randomization analyses were conducted, including continuous estradiol levels in the combined pre- and postmenopausal sample and BMI as exposure variables with brain age gap, Alzheimer’s disease, and depression as outcomes. Sensitivity analyses were conducted using the binary estradiol approach and, to avoid sample overlap, using the depression subsample excluding UKB. Furthermore, based on our univariable findings and following previous studies16,17,21, follow-up multivariable analyses were conducted, including age at menarche and BMI as exposures with depression, the depression subsample excluding UKB, and recurrent depression as outcomes. Using the TwoSampleMR package (v0.5.11)69, instrumental variables were extracted from the exposure datasets, the exposures and the outcome were harmonised, and multivariable IVW analyses were run. Conditional F-statistics for instrument strength were computed, and the multivariable MR-Egger method was applied to check the robustness of the multivariable Mendelian randomization results using the MVMR package (v0.1)72. The instrumental variables for the multivariable analyses with estradiol levels and BMI as exposures indicated no weak instrument bias for BMI, however, possible weak instrument bias for estradiol levels (brain age gap and Alzheimer’s disease as outcomes: conditional F = 1.92 for the continuous approach and 8.87 for the binary approach in the combined pre- and postmenopausal sample from the UKB; depression as an outcome: conditional F = 1.93 for the continuous approach and 9.16 for the binary approach in the combined pre- and postmenopausal sample from the UKB; depression subsample excluding UKB as an outcome: conditional F = 0.99 for the continuous approach and 8.49 for the binary approach in the combined pre- and postmenopausal sample from the UKB; see also Supplementary Note 3 and Supplementary Table 4). The instrumental variables included in the multivariable analyses of age at menarche and BMI as exposures did not exhibit weak instrument bias, except for possible weak instrument bias for BMI in the analysis with the depression subsample excluding UKB as an outcome (conditional F = 9.97).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link








:max_bytes(150000):strip_icc()/GettyImages-1644727881-73ef3c9679a6447faafc74eb0ff22f1e.jpg)
+ There are no comments
Add yours