Participant selection
All ethical regulations relevant to human research participants were followed. Each sample had their study procedures approved by the respective local ethics committees and obtained written informed consent by participants or their caregivers.
For training and cross validation, we used data obtained from the Philadelphia Neurodevelopmental Cohort (PNC)28 and from the Lifespan Human Connectome Project Development (HCP-D)29. After excluding 20 subjects in PNC and 8 subjects in HCP-D that were outliers for the Euler number, an image quality measure, we selected the same age range (8–22 years old) in both datasets, by excluding 19 subjects with an age range of 5 to 7 in the HCP-D. To correct for multiple scanners in image acquisition, we harmonized the volumes across the two dataset and the multiple scan sites in the HCP-D by using the neuroCombat harmonization for multi-site imaging data (version 1.0.13)60. The harmonization process was run across 5 scan sites (1 site for PNC, 4 site for HCP-D), taking into account sex and age expressed in months as covariates. To obtain a balanced sample for sex while controlling for age and brain size, we applied a matching procedure. We first split the participants according to their sex assigned at birth. Each female was then matched with one male according to the following criteria: 1) maximum absolute age difference of 12 months; 2) maximum absolute difference in the harmonized estimated total intracranial volume (eTIV) of 3%; 3) maximum absolute difference in Euler number of 1 standard deviation. If more than one male fitted the criteria, the one with the lower eTIV difference was selected as the best match. The final sample was composed of N = 1132 (50% females; PNC = 768 of which 48% females; HCP-D = 364 of which 53% females), with no significant difference in age (in months), eTIV or Euler number between sexes. Supplementary Table T1 shows the sample characteristic before and after the matching procedure for the selected and excluded participants.
To validate our models independently, we used data from two longitudinal studies: the Queensland Twin Adolescent Brain (QTAB)30 and the Adolescent Brain Cognitive Development (ABCD)33. The QTAB dataset includes adolescent twins (NBaseline = 392, 49% females, age at baseline: 9–14 years), scanned twice with an interval of 13 to 30 months (mean interval: 20 months). The ABCD study is a longitudinal study that follows participant throughout adolescence33. We used the data release 5.0, including two MRI scans collected 2 years apart, across multiple research study sites (NBaseline = 7792, 47% females, age at baseline: 9-11 years). After excluding 15 observations in QTAB and 279 observations in ABCD due to image quality issues, we harmonized the data across sites, using the LongCombat package (version 0.0.0.90000), the longitudinal ComBat harmonization for multi-scanner imaging data61. Supplementary Table T2 reports the included participants for each analysis.
Image segmentation and features selection
Raw T1-weigthed MRI scans were preprocessed in FreeSurfer (v. 7.1.1) and the cortical and subcortical reconstruction was carried out for all cross-sectional and longitudinal datasets. Although a longitudinal pipeline is available in FreeSurfer, this approach is discouraged when considering data from children, as it assumes a constant eTIV between sessions. We decided, therefore, to use the traditional pipeline, and account for the longitudinal setting in the harmonization process, as discussed in the previous section. A mix of defaced and non-defaced data was used for both the training (PNC not defaced, HCP-D defaced) and the two independent test samples (QTAB defaced, ABCD not defaced), as provided by the study sources. While defacing may impact brain measures, alignment of results across defaced and not defaced samples supports that findings are robust to potential defacing related issues.
As quality control of the imaging data, we used the Euler number, a proxy of image data quality35, averaged across the two hemispheres and excluded subjects with values lower than three standard deviations from the mean. The Euler number represents a measure of the topological complexity of the reconstructed cortex and has been shown to provide a useful proxy of structural image quality, given high association with manual rating procedures and outperforming motion indicators in identifying unusable images35,62. By excluding outliers for Euler number in each sample we aim to control for MRI quality reconstruction and potential motion artefacts.
For a better fine-tuning and more precise selection of limbic structures, we combined volumes derived from different segmentation approaches, as previously described elsewhere27. Briefly, we used the multimodal parcellation of the cerebral cortex63 together with multiple subcortical segmentation, including structures from the classical subcortical segmentation from FreeSurfer, the segmentation of subcortical limbic structures64 and the subfield segmentation of the hippocampus65, amygdala66 and thalamus64. The final feature set comprises 493 features (whole brain segmentation), carefully combined to avoid overlap between features derived with different segmentations. Each feature was then assigned to a limbic (160 features) or non-limbic (333 features) feature set based on the literature definition of the limbic system64,67,68,69. A complete list of limbic features and respective parcellation is listed in the Supplementary table T3, while Supplementary Fig. S13 illustrates the cortical distribution of limbic and non-limbic features.
To correct for differences in head size that could affect the model performance, prior to model training and external validation, we regressed out the eTIV from each feature and took the residuals.
Sex classification model
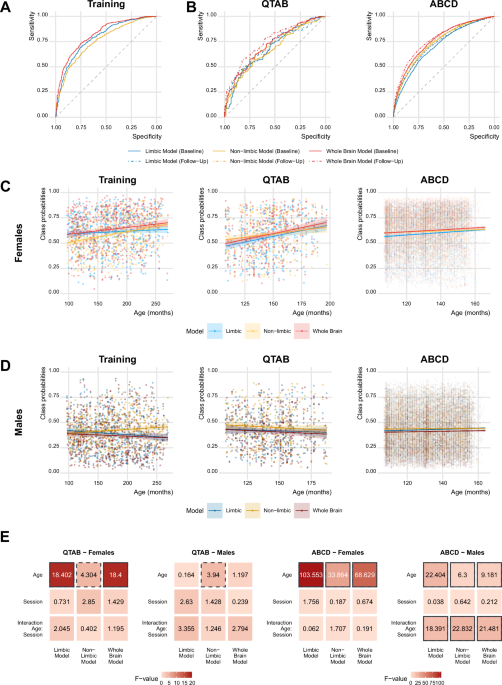
We trained one model for each set of features obtained after eTIV regression, using the binary classification implemented in the xgboost package (version 1.7.5.1) in R, following routines previously described in Matte Bon et al.27. In the training sample, sex was coded as a binary variable (males = 0, females = 1). Briefly, we performed a nested 5-fold cross validation, with the learning rate η = 0.01 and the initial number of rounds set to 1000. The prediction error in the inner loop was assessed for each iteration and used to determine the optimal number of iterations to train the final models on the entire set of data. The obtained class probabilities had a range between 0 and 1, where 0 corresponded to a male-like brain and 1 to a female-like brain. To assess each model’s performance, we computed the area under the receiving operating characteristic curves (AUC-ROC curves). To assess the external validation of our models in independent unseen data, we applied the trained models to the QTAB and ABCD data and stored the resulting class probabilities for further analyses. Although the QTAB dataset included genetically related participants (twins and siblings), the independence of the training sample ensured independent sex predictions in the validation samples.
Statistics and reproducibility
Biological associations with age and pubertal development
Since the effects of both age and pubertal development are expected to have opposite direction in males and females (i.e. positive associations in the direction of greater class probabilities in females towards more female-like brains, while negative associations in the direction of lower class probabilities in males toward more male-like brains), and a significant sex difference is expected in the class probabilities due to the classification model itself, all the analyses were carried out separately for each sex. For each set of analysis, three associations were run for each dataset, one for each machine learning model (limbic, non-limbic and whole brain).
The association with age was investigated in both the training and the two independent test samples. In the training data, a linear model for the class probabilities derived from each of the feature set was implemented, as follows:
$$subject( \, subject \sim _++).$$
In QTAB and ABCD, linear mixed effects (LME) models were run, to take into account the longitudinal nature of the data. For this we used the lme function of the nlme (version 3.1-165) package available in R, including age (expressed in months), session and the interaction between the two as the main independent variables, and to further control our analysis for Euler number. To capture the effects due to repeated measures, we included the subject as random effects in the model. The final model was written as follow:
$$subject\,( \sim \, _+subject+ \\ +subject_{months}:{session},{random}= \sim 1{|subject})$$
Since the ABCD has multiple study sites, we included the site as further covariate, after excluding sites with less than 20 observations available.
The associations with pubertal development were investigated using the QTAB and ABCD data. To do so, we used the average Youth Self-Report Pubertal Development Scale (PDS) score. This questionnaire consists of 5 questions, two of which are sex-specific, rating different aspects of pubertal development. Three common questions rate the growth in height, the body hair and change in skin. The two sex-specific questions refer in males to facial hair growth and voice break, while in females they refer to breast development and the beginning of menstruation. Each question is rated with a score from 1 to 4 as following: 1 = “Has not yet started”, 2 = “Has barely started”, 3 = “Is definitely underway” and 4 = “It is completed”. The question on menstruation in females constitutes the only exception and it is rated as binary, with “No” = 1 and “Yes” = 4. A caregiver report was also available and was used as further control for supplementary analyses in the ABCD sample.
We first assessed the direct association of class probabilities with the PDS score. To allow the direct comparisons of result between females and males, for each participant the average score of PDS question was calculated at each session and used as variable for the LME models. Each of the LME models for each dataset and set of features included the average PDS score, age (expressed in months) and the interaction between the two, as well as the Euler number as further covariates. Since the PDS average score is expected to increase continuously with age independently from the study session, session was not included in this analysis. The possible correlation between observations due to the longitudinal nature of the data was captured adding a random effect due to subject, as follows:
$${lme}({Class}{Probabilities} \sim \, {{PDS}}_{{average}}+{{age}}_{{months}}+{Euler} \\ +{{PDS}}_{{average}}:{{age}}_{{months}},{random}= \sim 1{|subject}).$$
For the ABCD, site was included as a further covariate, after excluding sites with less than 20 observations available.
We applied an ANOVA type I to the output of each LME model, to investigate the main effect of PDS score. As further control and to disentangle the relative contributions of the covariates, we ran an ANOVA type II. The results of these analyses are presented in the Supplementary File.
Next, we assessed the association with menarche onset in both datasets. For this, we divided the females into two groups based on their answers to the PDS question about menarche: 1- females subjects that did not start to menstruate at any of the two sessions of the studies (answered 1 for both session the relative PDS question); 2- females that had menarche (i.e. first menstrual cycle) between the two sessions (answered 1 at baseline and 4 at follow-up). The final sample for QTAB included a total of N = 110 participants at baseline (of which n = 46 participants had menarche between the two sessions), and N = 1091 at baseline for ABCD (of which n = 414 participants had menarche between sessions). We then ran an LME model for each machine learning model, similarly to the one described for PDS score, including age (expressed in months), session, the interaction between age and menarche, the interaction between session and menarche and Euler number, capturing the random effect due to repeated measure in the same subject. In the ABCD, we further added as covariate the site, after removing sites with less than 20 observations. The final model was written as follows:
$${lme} \, ({Class}\,{Probabilities} \sim {Menarche}\,{onset}+{{age}}_{{months}}+{session}+{Euler}+{menarche}\,{onset} \, {{age}}_{{months}} \\ +{menarche}\,{onset}:{session},{random}= \, \sim \, 1{|subject})$$
We applied an ANOVA type I to the output of LME to assess the main effect of menarche onset. We further tested the relative contributions of menarche, age and session by applying an ANOVA type II (Supplementary File).
Mental health associations
To investigate the association with mental health in the QTAB sample, we used data from the Spence Children’s Anxiety Scale (SCAS), the Short Mood and Feeling Questionnaire (SMFQ) and the anxiety and depression subscale of the Somatic and Psychological Health Report (SPHERE-21) obtained from the self-reported questionnaires. These questionnaires measure symptoms of anxiety and depression and were collected at both sessions, allowing us to assess longitudinal changes in mental health scores. To obtain a unique continuous score, we ran a principal component analysis (PCA) for dimensionality reduction across all questions at baseline and took the first component as a general score for mental health. To assess the mental health score at follow-up, we took the eigenvectors of each question for the first component at baseline and applied them to the data at follow-up, obtaining the PCA scores.
To assess the association of mental health scores with class probabilities, we run an LME model for each machine learning model as following:
$${lme} \, ({Class}{Probabilities} \sim \, {PCA\; scores}+{{age}}_{{months}}+{session}+{Euler} +{PCA\; scores}: \\ {{age}}_{{months}}+{PCA\; scores}:{session},{random} \, = \, \sim 1{|subject})$$
We tested for significance of the main effect of mental health by applying an ANOVA type I to the LME output. Further, we tested relative contributions of covariates with an ANOVA type II (see Supplementary File for results).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link




:max_bytes(150000):strip_icc()/What-Happens-to-Your-Brain-When-You-Drink-A-Diet-Soda-Every-Day-3656a7a7704440af8ef3be355b5e77bd.jpg)



:max_bytes(150000):strip_icc()/GettyImages-1644727881-73ef3c9679a6447faafc74eb0ff22f1e.jpg)
+ There are no comments
Add yours